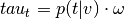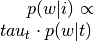nimare.decode.continuous.gclda_decode_map
- gclda_decode_map(model, image, topic_priors=None, prior_weight=1)[source]
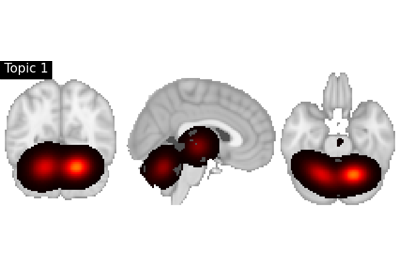
Perform image-to-text decoding for continuous inputs using method from Rubin et al. (2017).
- Parameters
model (
nimare.annotate.topic.GCLDAModel) – Model object needed for decoding.image (
nibabel.nifti1.Nifti1Imageorstr) – Whole-brain image to decode into text. Must be in same space as model and dataset. Model’s template available in model.dataset.mask_img.topic_priors (
numpy.ndarrayoffloat, optional) – A 1d array of size (n_topics) with values for topic weighting. If None, no weighting is done. Default is None.prior_weight (
float, optional) – The weight by which the prior will affect the decoding. Default is 1.
- Returns
decoded_df (
pandas.DataFrame) – A DataFrame with the word-tokens and their associated weights.topic_weights (
numpy.ndarrayoffloat) – The weights of the topics used in decoding.
Notes
Notation
Meaning

Voxel
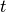
Topic
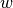
Word type
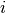
Input image
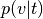
Probability of topic given voxel (
p_topic_g_voxel)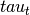
Topic weight vector (
topic_weights)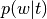
Probability of word type given topic (
p_word_g_topic)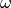
1d array from input image (
input_values)Compute
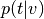 (
(p_topic_g_voxel).From
gclda.model.Model.get_spatial_probs()
Squeeze input image to 1d array
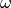 (
(input_values).Compute topic weight vector (
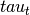 ) by multiplying
) by multiplying
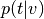 by input image.
by input image.Multiply
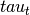 by
by
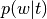 .
.The resulting vector (
word_weights) reflects arbitrarily scaled term weights for the input image.
See also
nimare.annotate.gclda.GCLDAModel,nimare.decode.discrete.gclda_decode_roi(),nimare.decode.encode.gclda_encode()References
Rubin, Timothy N., et al. “Decoding brain activity using a large-scale probabilistic functional-anatomical atlas of human cognition.” PLoS computational biology 13.10 (2017): e1005649. https://doi.org/10.1371/journal.pcbi.1005649