Note
Click here to download the full example code
Train a GCLDA model and use it
This example trains a generalized correspondence latent Dirichlet allocation model using abstracts from Neurosynth and then uses it for decoding.
Warning
The model in this example is trained using (1) a very small, nonrepresentative dataset and (2) very few iterations. As such, it will not provide useful results. If you are interested in using GCLDA, we recommend using a large dataset like Neurosynth, and training with at least 10k iterations.
import os
import nibabel as nib
import numpy as np
from nilearn import image, masking, plotting
import nimare
from nimare import annotate, decode
from nimare.tests.utils import get_test_data_path
Load dataset with abstracts
We’ll load a small dataset composed only of studies in Neurosynth with Angela Laird as a coauthor, for the sake of speed.
dset = nimare.dataset.Dataset(os.path.join(get_test_data_path(), "neurosynth_laird_studies.json"))
dset.texts.head(2)
Generate term counts
GCLDA uses raw word counts instead of the tf-idf values generated by Neurosynth.
counts_df = annotate.text.generate_counts(
dset.texts,
text_column="abstract",
tfidf=False,
max_df=0.99,
min_df=0.01,
)
counts_df.head(5)
Out:
/home/docs/checkouts/readthedocs.org/user_builds/nimare/envs/0.0.10/lib/python3.7/site-packages/sklearn/utils/deprecation.py:87: FutureWarning: Function get_feature_names is deprecated; get_feature_names is deprecated in 1.0 and will be removed in 1.2. Please use get_feature_names_out instead.
warnings.warn(msg, category=FutureWarning)
Run model
Five iterations will take ~10 minutes with the full Neurosynth dataset. It’s much faster with this reduced example dataset. Note that we’re using only 10 topics here. This is because there are only 13 studies in the dataset. If the number of topics is higher than the number of studies in the dataset, errors can occur during training.
model = annotate.gclda.GCLDAModel(
counts_df,
dset.coordinates,
mask=dset.masker.mask_img,
n_topics=10,
n_regions=4,
symmetric=True,
)
model.fit(n_iters=100, loglikely_freq=20)
model.save("gclda_model.pkl.gz")
# Let's remove the model now that you know how to generate it.
os.remove("gclda_model.pkl.gz")
Out:
/home/docs/checkouts/readthedocs.org/user_builds/nimare/checkouts/0.0.10/nimare/annotate/gclda.py:180: FutureWarning: In a future version of pandas all arguments of DataFrame.drop except for the argument 'labels' will be keyword-only
count_df = count_df.drop("id", 1)
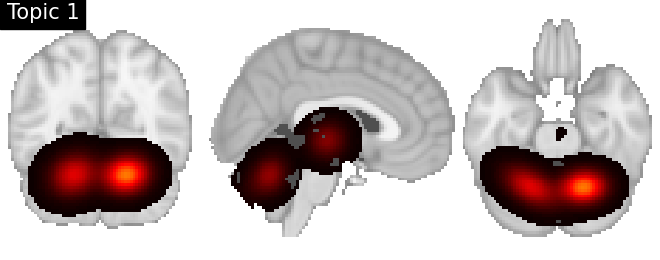
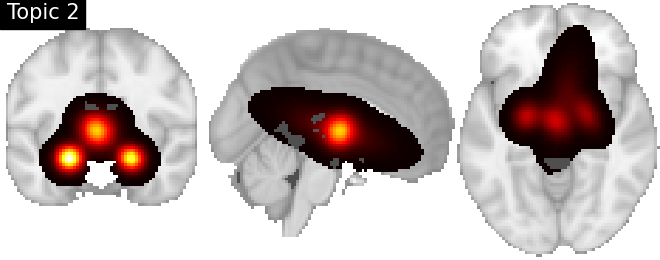
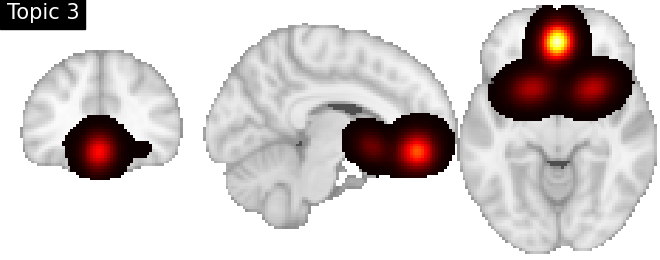
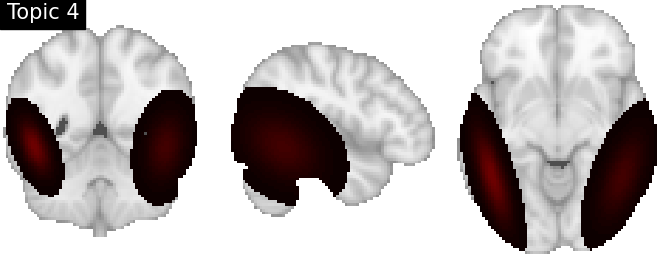
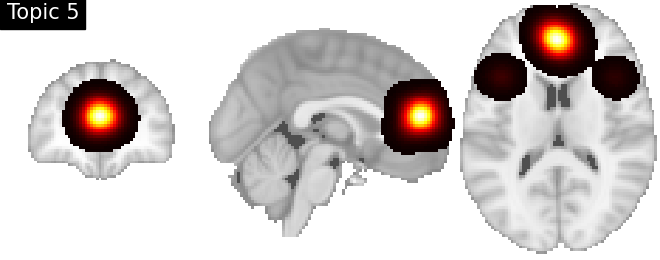
Look at topics
topic_img_4d = masking.unmask(model.p_voxel_g_topic_.T, model.mask)
for i_topic in range(5):
topic_img_3d = image.index_img(topic_img_4d, i_topic)
plotting.plot_stat_map(
topic_img_3d,
draw_cross=False,
colorbar=False,
annotate=False,
title=f"Topic {i_topic + 1}",
)
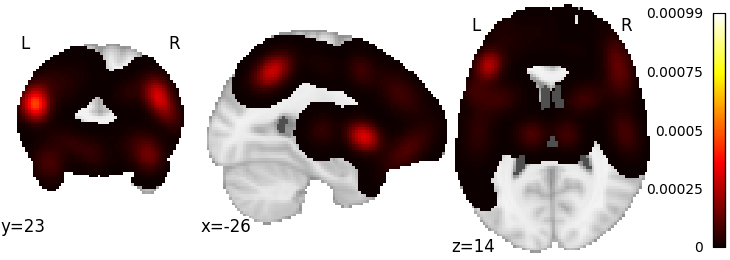
Generate a pseudo-statistic image from text
text = "dorsal anterior cingulate cortex"
encoded_img, _ = decode.encode.gclda_encode(model, text)
plotting.plot_stat_map(encoded_img, draw_cross=False)
Out:
<nilearn.plotting.displays.OrthoSlicer object at 0x7f1889abba50>
Decode an unthresholded statistical map
For the sake of simplicity, we will use the pseudo-statistic map generated in the previous step.
# Run the decoder
decoded_df, _ = decode.continuous.gclda_decode_map(model, encoded_img)
decoded_df.sort_values(by="Weight", ascending=False).head(10)
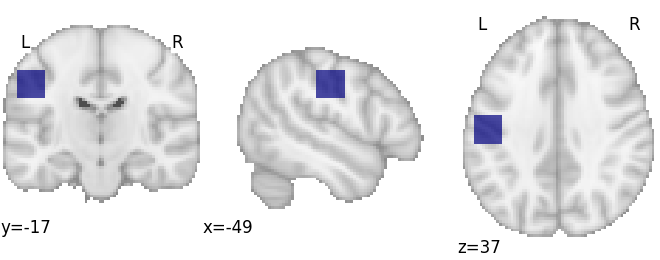
Decode an ROI image
First we’ll make an ROI
arr = np.zeros(dset.masker.mask_img.shape, int)
arr[65:75, 50:60, 50:60] = 1
mask_img = nib.Nifti1Image(arr, dset.masker.mask_img.affine)
plotting.plot_roi(mask_img, draw_cross=False)
Out:
<nilearn.plotting.displays.OrthoSlicer object at 0x7f1889a7f190>
Run the decoder
decoded_df, _ = decode.discrete.gclda_decode_roi(model, mask_img)
decoded_df.sort_values(by="Weight", ascending=False).head(10)
Total running time of the script: ( 0 minutes 41.164 seconds)