Note
Click here to download the full example code
Test combinations of kernels and estimators for coordinate-based meta-analyses.
Collection of NIDM-Results packs downloaded from Neurovault collection 1425, uploaded by Dr. Camille Maumet.
Note
Creation of the Dataset from the NIDM-Results packs was done with custom code. The Results packs for collection 1425 are not completely NIDM-Results-compliant, so the nidmresults library could not be used to facilitate data extraction.
import os
from nilearn.plotting import plot_stat_map
import nimare
from nimare.tests.utils import get_test_data_path
Load Dataset
dset_file = os.path.join(get_test_data_path(), "nidm_pain_dset.json")
dset = nimare.dataset.Dataset(dset_file)
mask_img = dset.masker.mask_img
List possible kernel transformers
kernel_transformers = {
"MKDA kernel": nimare.meta.kernel.MKDAKernel,
"KDA kernel": nimare.meta.kernel.KDAKernel,
"ALE kernel": nimare.meta.kernel.ALEKernel,
}
MKDA density analysis
for kt_name, kt in kernel_transformers.items():
try:
mkda = nimare.meta.MKDADensity(kernel_transformer=kt, null_method="approximate")
mkda.fit(dset)
corr = nimare.correct.FWECorrector(method="montecarlo", n_iters=10, n_cores=1)
cres = corr.transform(mkda.results)
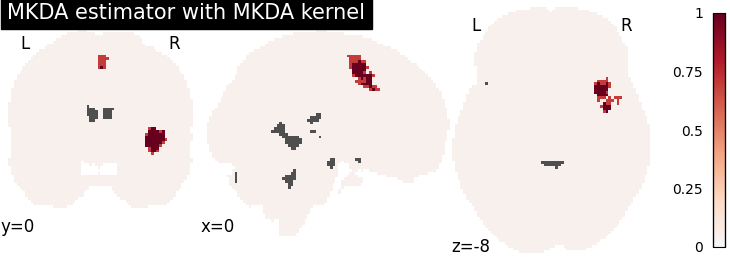
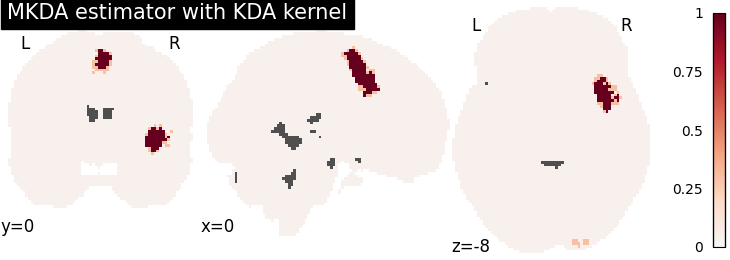
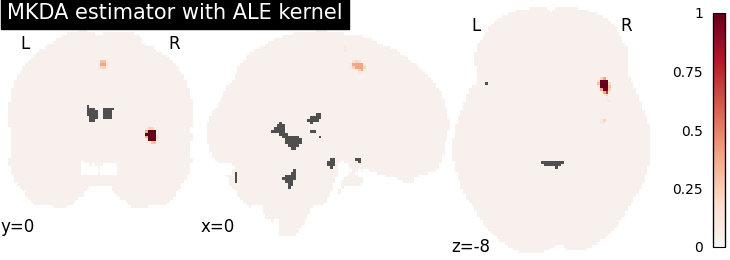
plot_stat_map(
cres.get_map("logp_level-voxel_corr-FWE_method-montecarlo"),
cut_coords=[0, 0, -8],
draw_cross=False,
cmap="RdBu_r",
title="MKDA estimator with %s" % kt_name,
)
except AttributeError:
print(
"\nError: the %s does not currently work with the MKDA meta-analysis method\n"
% kt_name
)
Out:
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:02, 3.33it/s]
20%|## | 2/10 [00:00<00:02, 3.34it/s]
30%|### | 3/10 [00:00<00:02, 3.34it/s]
40%|#### | 4/10 [00:01<00:01, 3.34it/s]
50%|##### | 5/10 [00:01<00:01, 3.35it/s]
60%|###### | 6/10 [00:01<00:01, 3.34it/s]
70%|####### | 7/10 [00:02<00:00, 3.35it/s]
80%|######## | 8/10 [00:02<00:00, 3.35it/s]
90%|######### | 9/10 [00:02<00:00, 3.35it/s]
100%|##########| 10/10 [00:02<00:00, 3.35it/s]
100%|##########| 10/10 [00:02<00:00, 3.34it/s]
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:02, 3.12it/s]
20%|## | 2/10 [00:00<00:02, 3.13it/s]
30%|### | 3/10 [00:00<00:02, 3.13it/s]
40%|#### | 4/10 [00:01<00:01, 3.10it/s]
50%|##### | 5/10 [00:01<00:01, 3.12it/s]
60%|###### | 6/10 [00:01<00:01, 3.13it/s]
70%|####### | 7/10 [00:02<00:00, 3.14it/s]
80%|######## | 8/10 [00:02<00:00, 3.14it/s]
90%|######### | 9/10 [00:02<00:00, 3.14it/s]
100%|##########| 10/10 [00:03<00:00, 3.15it/s]
100%|##########| 10/10 [00:03<00:00, 3.14it/s]
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:03, 2.69it/s]
20%|## | 2/10 [00:00<00:02, 2.69it/s]
30%|### | 3/10 [00:01<00:02, 2.68it/s]
40%|#### | 4/10 [00:01<00:02, 2.68it/s]
50%|##### | 5/10 [00:01<00:01, 2.68it/s]
60%|###### | 6/10 [00:02<00:01, 2.69it/s]
70%|####### | 7/10 [00:02<00:01, 2.69it/s]
80%|######## | 8/10 [00:02<00:00, 2.69it/s]
90%|######### | 9/10 [00:03<00:00, 2.69it/s]
100%|##########| 10/10 [00:03<00:00, 2.70it/s]
100%|##########| 10/10 [00:03<00:00, 2.69it/s]
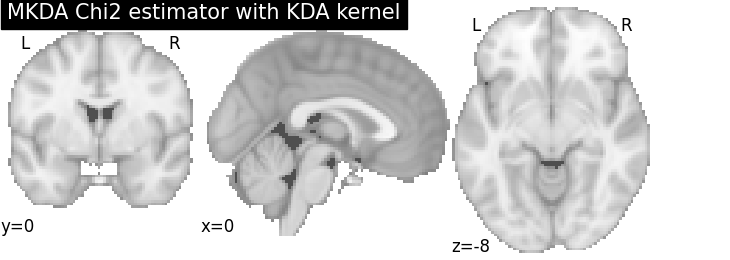
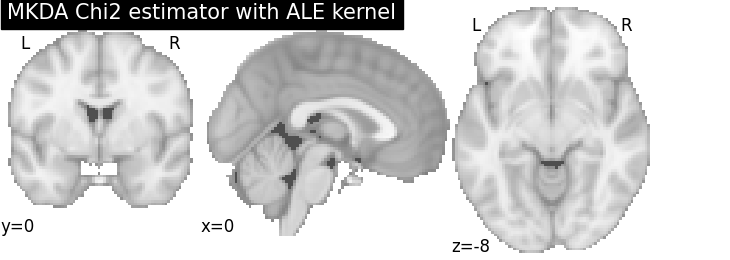
MKDA Chi2
for kt_name, kt in kernel_transformers.items():
try:
mkda = nimare.meta.MKDAChi2(kernel_transformer=kt)
dset1 = dset.slice(dset.ids)
dset2 = dset.slice(dset.ids)
mkda.fit(dset1, dset2)
corr = nimare.correct.FWECorrector(method="montecarlo", n_iters=10, n_cores=1)
cres = corr.transform(mkda.results)
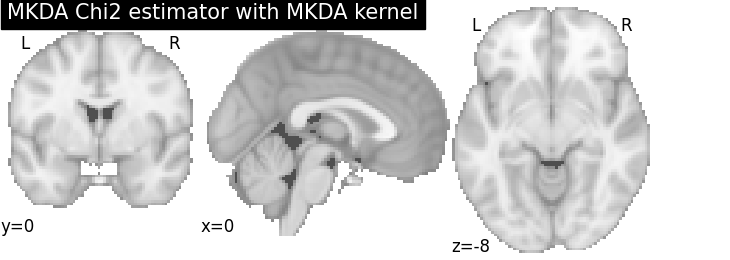
plot_stat_map(
cres.get_map("z_desc-consistency_level-voxel_corr-FWE_method-montecarlo"),
threshold=1.65,
cut_coords=[0, 0, -8],
draw_cross=False,
cmap="RdBu_r",
title="MKDA Chi2 estimator with %s" % kt_name,
)
except AttributeError:
print(
"\nError: the %s does not currently work with the MKDA Chi2 meta-analysis method\n"
% kt_name
)
Out:
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:03, 2.78it/s]
20%|## | 2/10 [00:00<00:02, 2.83it/s]
30%|### | 3/10 [00:01<00:02, 2.86it/s]
40%|#### | 4/10 [00:01<00:02, 2.86it/s]
50%|##### | 5/10 [00:01<00:01, 2.86it/s]
60%|###### | 6/10 [00:02<00:01, 2.87it/s]
70%|####### | 7/10 [00:02<00:01, 2.87it/s]
80%|######## | 8/10 [00:02<00:00, 2.87it/s]
90%|######### | 9/10 [00:03<00:00, 2.87it/s]
100%|##########| 10/10 [00:03<00:00, 2.87it/s]
100%|##########| 10/10 [00:03<00:00, 2.86it/s]
/home/docs/checkouts/readthedocs.org/user_builds/nimare/envs/0.0.11rc1/lib/python3.7/site-packages/nilearn/plotting/displays.py:880: UserWarning: empty mask
get_mask_bounds(new_img_like(img, not_mask, affine))
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:03, 2.54it/s]
20%|## | 2/10 [00:00<00:03, 2.57it/s]
30%|### | 3/10 [00:01<00:02, 2.59it/s]
40%|#### | 4/10 [00:01<00:02, 2.59it/s]
50%|##### | 5/10 [00:01<00:01, 2.60it/s]
60%|###### | 6/10 [00:02<00:01, 2.60it/s]
70%|####### | 7/10 [00:02<00:01, 2.60it/s]
80%|######## | 8/10 [00:03<00:00, 2.61it/s]
90%|######### | 9/10 [00:03<00:00, 2.61it/s]
100%|##########| 10/10 [00:03<00:00, 2.61it/s]
100%|##########| 10/10 [00:03<00:00, 2.60it/s]
/home/docs/checkouts/readthedocs.org/user_builds/nimare/envs/0.0.11rc1/lib/python3.7/site-packages/nilearn/plotting/displays.py:880: UserWarning: empty mask
get_mask_bounds(new_img_like(img, not_mask, affine))
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:04, 2.00it/s]
20%|## | 2/10 [00:00<00:03, 2.02it/s]
30%|### | 3/10 [00:01<00:03, 2.02it/s]
40%|#### | 4/10 [00:01<00:02, 2.02it/s]
50%|##### | 5/10 [00:02<00:02, 2.02it/s]
60%|###### | 6/10 [00:02<00:01, 2.03it/s]
70%|####### | 7/10 [00:03<00:01, 2.03it/s]
80%|######## | 8/10 [00:03<00:00, 2.03it/s]
90%|######### | 9/10 [00:04<00:00, 2.03it/s]
100%|##########| 10/10 [00:04<00:00, 2.03it/s]
100%|##########| 10/10 [00:04<00:00, 2.02it/s]
/home/docs/checkouts/readthedocs.org/user_builds/nimare/envs/0.0.11rc1/lib/python3.7/site-packages/nilearn/plotting/displays.py:880: UserWarning: empty mask
get_mask_bounds(new_img_like(img, not_mask, affine))
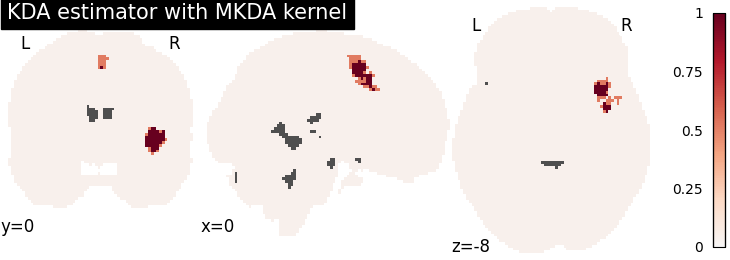
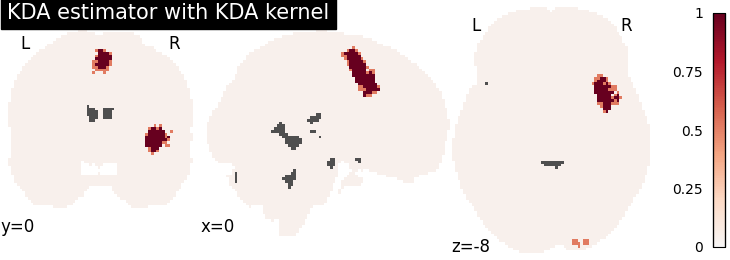
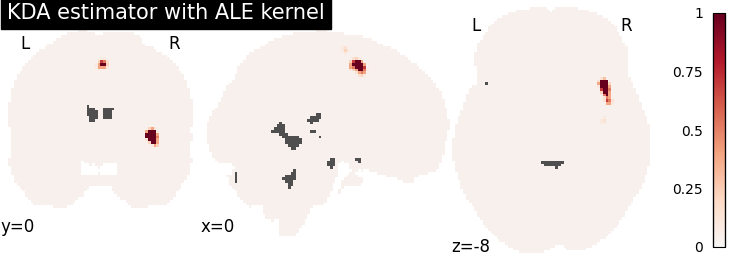
KDA
for kt_name, kt in kernel_transformers.items():
try:
kda = nimare.meta.KDA(kernel_transformer=kt, null_method="approximate")
kda.fit(dset)
corr = nimare.correct.FWECorrector(method="montecarlo", n_iters=10, n_cores=1)
cres = corr.transform(kda.results)
plot_stat_map(
cres.get_map("logp_level-voxel_corr-FWE_method-montecarlo"),
cut_coords=[0, 0, -8],
draw_cross=False,
cmap="RdBu_r",
title="KDA estimator with %s" % kt_name,
)
except AttributeError:
print(
"\nError: the %s does not currently work with the KDA meta-analysis method\n" % kt_name
)
Out:
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:02, 3.50it/s]
20%|## | 2/10 [00:00<00:02, 3.57it/s]
30%|### | 3/10 [00:00<00:01, 3.61it/s]
40%|#### | 4/10 [00:01<00:01, 3.62it/s]
50%|##### | 5/10 [00:01<00:01, 3.63it/s]
60%|###### | 6/10 [00:01<00:01, 3.65it/s]
70%|####### | 7/10 [00:01<00:00, 3.66it/s]
80%|######## | 8/10 [00:02<00:00, 3.65it/s]
90%|######### | 9/10 [00:02<00:00, 3.67it/s]
100%|##########| 10/10 [00:02<00:00, 3.67it/s]
100%|##########| 10/10 [00:02<00:00, 3.64it/s]
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:02, 3.37it/s]
20%|## | 2/10 [00:00<00:02, 3.38it/s]
30%|### | 3/10 [00:00<00:02, 3.38it/s]
40%|#### | 4/10 [00:01<00:01, 3.37it/s]
50%|##### | 5/10 [00:01<00:01, 3.38it/s]
60%|###### | 6/10 [00:01<00:01, 3.39it/s]
70%|####### | 7/10 [00:02<00:00, 3.38it/s]
80%|######## | 8/10 [00:02<00:00, 3.39it/s]
90%|######### | 9/10 [00:02<00:00, 3.39it/s]
100%|##########| 10/10 [00:02<00:00, 3.39it/s]
100%|##########| 10/10 [00:02<00:00, 3.38it/s]
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:03, 2.99it/s]
20%|## | 2/10 [00:00<00:02, 2.99it/s]
30%|### | 3/10 [00:01<00:02, 2.99it/s]
40%|#### | 4/10 [00:01<00:02, 2.99it/s]
50%|##### | 5/10 [00:01<00:01, 2.98it/s]
60%|###### | 6/10 [00:02<00:01, 2.98it/s]
70%|####### | 7/10 [00:02<00:01, 2.98it/s]
80%|######## | 8/10 [00:02<00:00, 2.98it/s]
90%|######### | 9/10 [00:03<00:00, 2.97it/s]
100%|##########| 10/10 [00:03<00:00, 2.98it/s]
100%|##########| 10/10 [00:03<00:00, 2.98it/s]
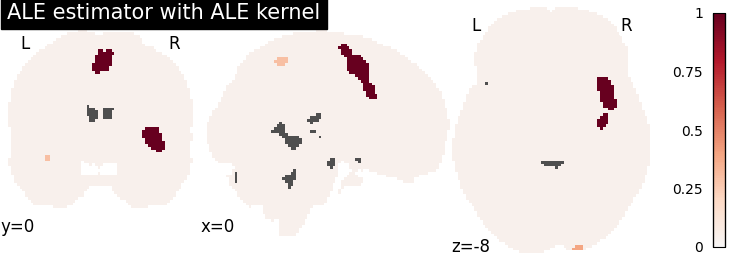
ALE
for kt_name, kt in kernel_transformers.items():
try:
ale = nimare.meta.ALE(kernel_transformer=kt, null_method="approximate")
ale.fit(dset)
corr = nimare.correct.FWECorrector(method="montecarlo", n_iters=10, n_cores=1)
cres = corr.transform(ale.results)
plot_stat_map(
cres.get_map("logp_level-cluster_corr-FWE_method-montecarlo"),
cut_coords=[0, 0, -8],
draw_cross=False,
cmap="RdBu_r",
title="ALE estimator with %s" % kt_name,
)
except AttributeError:
print(
"\nError: the %s does not currently work with the ALE meta-analysis method\n" % kt_name
)
Out:
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:02, 3.16it/s]
20%|## | 2/10 [00:00<00:02, 3.28it/s]
30%|### | 3/10 [00:00<00:02, 3.30it/s]
40%|#### | 4/10 [00:01<00:01, 3.32it/s]
50%|##### | 5/10 [00:01<00:01, 3.31it/s]
60%|###### | 6/10 [00:01<00:01, 3.32it/s]
70%|####### | 7/10 [00:02<00:00, 3.33it/s]
80%|######## | 8/10 [00:02<00:00, 3.33it/s]
90%|######### | 9/10 [00:02<00:00, 3.34it/s]
100%|##########| 10/10 [00:03<00:00, 3.33it/s]
100%|##########| 10/10 [00:03<00:00, 3.32it/s]
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:02, 3.12it/s]
20%|## | 2/10 [00:00<00:02, 3.11it/s]
30%|### | 3/10 [00:00<00:02, 3.13it/s]
40%|#### | 4/10 [00:01<00:01, 3.12it/s]
50%|##### | 5/10 [00:01<00:01, 3.12it/s]
60%|###### | 6/10 [00:01<00:01, 3.12it/s]
70%|####### | 7/10 [00:02<00:00, 3.13it/s]
80%|######## | 8/10 [00:02<00:00, 3.14it/s]
90%|######### | 9/10 [00:02<00:00, 3.14it/s]
100%|##########| 10/10 [00:03<00:00, 3.13it/s]
100%|##########| 10/10 [00:03<00:00, 3.13it/s]
0%| | 0/10 [00:00<?, ?it/s]
10%|# | 1/10 [00:00<00:03, 2.85it/s]
20%|## | 2/10 [00:00<00:02, 2.87it/s]
30%|### | 3/10 [00:01<00:02, 2.88it/s]
40%|#### | 4/10 [00:01<00:02, 2.88it/s]
50%|##### | 5/10 [00:01<00:01, 2.88it/s]
60%|###### | 6/10 [00:02<00:01, 2.88it/s]
70%|####### | 7/10 [00:02<00:01, 2.87it/s]
80%|######## | 8/10 [00:02<00:00, 2.87it/s]
90%|######### | 9/10 [00:03<00:00, 2.88it/s]
100%|##########| 10/10 [00:03<00:00, 2.88it/s]
100%|##########| 10/10 [00:03<00:00, 2.87it/s]
Total running time of the script: ( 1 minutes 43.879 seconds)