Note
Click here to download the full example code
Compare image and coordinate based meta-analyses
Run IBMAs and CBMAs on a toy dataset, then compare the results qualitatively.
Collection of NIDM-Results packs downloaded from Neurovault collection 1425, uploaded by Dr. Camille Maumet.
import os
import pandas as pd
from nilearn.plotting import plot_stat_map
from nimare.dataset import Dataset
from nimare.extract import download_nidm_pain
from nimare.meta.cbma import ALE
from nimare.meta.ibma import DerSimonianLaird
from nimare.transforms import ImagesToCoordinates, ImageTransformer
from nimare.utils import get_resource_path
Download data
Load Dataset
dset_file = os.path.join(get_resource_path(), "nidm_pain_dset.json")
dset = Dataset(dset_file)
dset.update_path(dset_dir)
# Calculate missing statistical images from the available stats.
xformer = ImageTransformer(target=["z", "varcope"])
dset = xformer.transform(dset)
# create coordinates from statistical maps
coord_gen = ImagesToCoordinates(merge_strategy="replace")
dset = coord_gen.transform(dset)
ALE (CBMA)
meta_cbma = ALE()
meta_cbma.fit(dset)
plot_stat_map(
meta_cbma.results.get_map("z"), cut_coords=[0, 0, -8], draw_cross=False, cmap="RdBu_r"
)
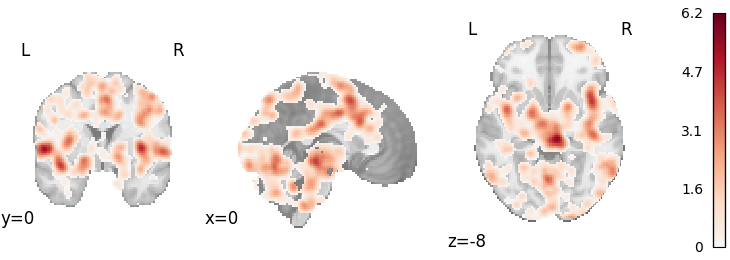
Out:
<nilearn.plotting.displays._slicers.OrthoSlicer object at 0x7fc437ee75d0>
DerSimonian-Laird (IBMA)
We must resample the image data to the same MNI template as the Dataset.
meta_ibma = DerSimonianLaird(resample=True)
meta_ibma.fit(dset)
plot_stat_map(
meta_ibma.results.get_map("z"), cut_coords=[0, 0, -8], draw_cross=False, cmap="RdBu_r"
)
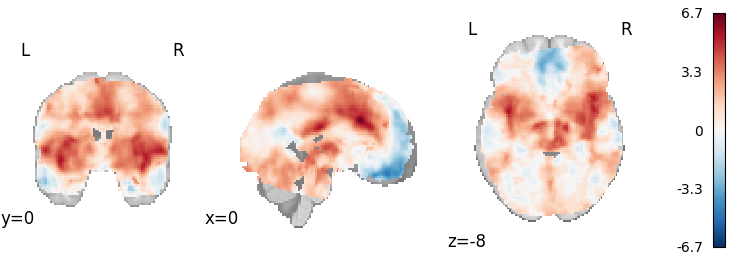
Out:
/home/docs/checkouts/readthedocs.org/user_builds/nimare/envs/0.0.12rc1/lib/python3.7/site-packages/nilearn/_utils/niimg.py:62: UserWarning: Non-finite values detected. These values will be replaced with zeros.
"Non-finite values detected. "
<nilearn.plotting.displays._slicers.OrthoSlicer object at 0x7fc437dc3a90>
Compare CBMA and IBMA Z-maps
stat_df = pd.DataFrame(
{
"CBMA": meta_cbma.results.get_map("z", return_type="array"),
"IBMA": meta_ibma.results.get_map("z", return_type="array").squeeze(),
}
)
print(stat_df.corr())
Out:
CBMA IBMA
CBMA 1.000000 0.516817
IBMA 0.516817 1.000000
Total running time of the script: ( 4 minutes 11.315 seconds)