Note
Click here to download the full example code
Using Neurovault Statistical Maps in NiMARE¶
Conversion of Statistical Maps¶
Some of the statistical maps are T statistics and others are Z statistics.
To perform a Fisher’s meta analysis, we need all Z maps.
Thoughtfully, NiMARE has a function named transform_images that will
help us.
from nimare.transforms import transform_images
# Not all studies have Z maps!
print(dset.images["z"])
dset.images = transform_images(
dset.images, target="z", masker=dset.masker, metadata_df=dset.metadata
)
# All studies have Z maps!
print(dset.images["z"])
Out:
1 None
0 None
2 None
5 /home/docs/.nimare/working_memory/collection-3...
4 /home/docs/.nimare/working_memory/collection-3...
6 /home/docs/.nimare/working_memory/collection-4...
3 None
Name: z, dtype: object
/home/docs/checkouts/readthedocs.org/user_builds/nimare/envs/0.0.8/lib/python3.7/site-packages/nilearn/image/resampling.py:598: RuntimeWarning: NaNs or infinite values are present in the data passed to resample. This is a bad thing as they make resampling ill-defined and much slower.
fill_value=fill_value)
1 /home/docs/.nimare/working_memory/study-2621-w...
0 /home/docs/.nimare/working_memory/study-2884-w...
2 /home/docs/.nimare/working_memory/study-3085-w...
5 /home/docs/.nimare/working_memory/collection-3...
4 /home/docs/.nimare/working_memory/collection-3...
6 /home/docs/.nimare/working_memory/collection-4...
3 /home/docs/.nimare/working_memory/study-5623-w...
Name: z, dtype: object
Run a Meta-Analysis¶
With the missing Z maps filled in, we can run a Meta-Analysis and plot our results
from nimare.meta.ibma import Fishers
from nilearn.plotting import plot_stat_map
meta = Fishers()
meta_res = meta.fit(dset)
plot_stat_map(meta_res.get_map("z"), threshold=3.3)
# The result may look questionable, but this code provides
# a template on how to use neurovault in your meta analysis.
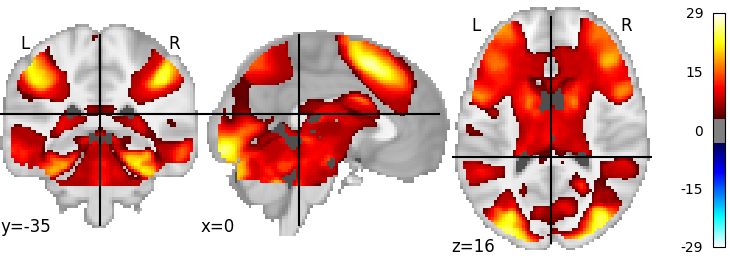
Out:
<nilearn.plotting.displays.OrthoSlicer object at 0x7f4fda984f90>
Total running time of the script: ( 0 minutes 16.731 seconds)