About NiMARE
NiMARE is a Python package for performing meta-analyses, and derivative analyses using meta-analytic data, of the neuroimaging literature. While meta-analytic packages exist which implement one or two algorithms each, NiMARE provides a standard syntax for performing a wide range of analyses and for interacting with databases of coordinates and images from fMRI studies (e.g., brainspell, Neurosynth, and NeuroVault).
NiMARE joins a growing Python ecosystem for neuroimaging research, which includes such tools as Nipype, Nibabel, and Nilearn. As with these other tools, NiMARE is open source, collaboratively developed, and built with ease of use in mind.
This page outlines NiMARE’s purpose and its role in a proposed meta-analytic ecosystem.
NiMARE’s Scope and Roadmap
NiMARE’s primary goal is to consolidate coordinate- and image-based meta-analysis methods with a simple, shared and comprehensive interface. This should reduce brand loyalty to any given algorithm, as it should be easy to employ the most appropriate algorithm for a given project. NiMARE also provides an environment where comparisons between methods are easier to perform.
A secondary goal of NiMARE is to implement some of the more cutting-edge methods for analyses built on meta-analytic neuroimaging data. There are many tools or algorithms that use meta-analytic data, including automated annotation, meta-analytic functional characterization analysis, and meta-analytic parcellation. Many of these methods are either tied to a specific meta-analysis package or never make it from publication to useable (i.e., documented and tested) code.
NiMARE’s Role in a Proposed Meta-Analytic Ecosystem
Important
For more up-to-date information and for information about other elements in the ecosystem, please see neurostuff.github.io.
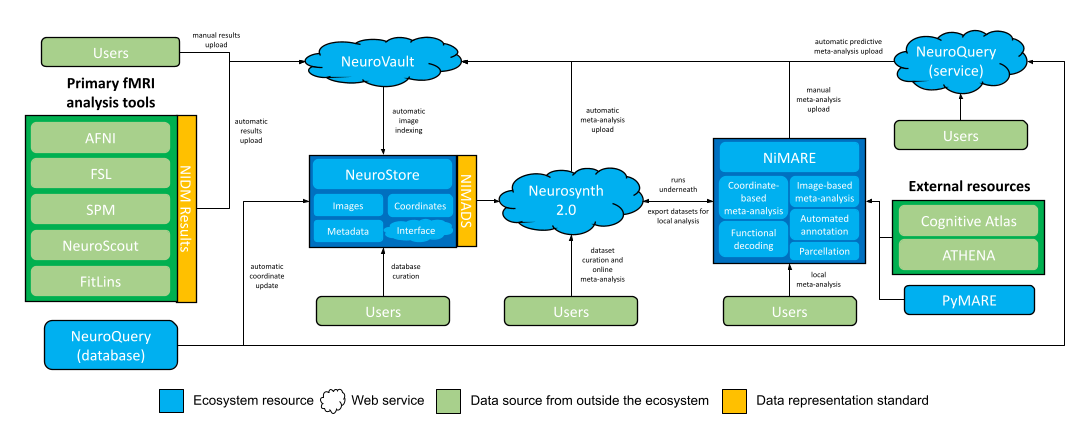
NiMARE aims to fill a gap in a burgeoning meta-analytic ecosystem. The goal of NiMARE is to collect a wide range of meta-analytic tools in one Python library. Currently, those methods are spread out across a range of programming languages and user interfaces, or are never even translated from the original papers into useable tools. NiMARE operates on NIMADS-format datasets, which users will be able to compile by searching the NeuroStore database with the pyNIMADS library. A number of other services in the ecosystem will then use NiMARE functions to perform meta-analyses, including Neurosynth 2.0 and NeuroVault.
Other Meta-Analytic Tools
Outside of the shared ecosystem detailed above, there are a number of tools.
Coordinate-based meta-analysis tools
BrainMap: The BrainMap suite includes applications for the ALE CBMA algorithm (via the GingerALE app) and interacting with the BrainMap database (via the Sleuth and Scribe apps).
The MKDA Toolbox: This toolbox implements the MKDA algorithm, as well as a range derivative analyses.
The SDM Toolbox: This toolbox contains the hybrid coordinate/image-based seed-based d-mapping algorithm.
NeuRoi Toolbox: This toolbox contains an implementation of the Analysis of Brain Coordinates (ABC) CBMA algorithm.
Image-based meta-analysis tools
IBMA SPM extension: This SPM extension implements a number of image-based meta-analysis algorithms.
Meta-analysis tools for other neuroimaging modalities
ERPscanr: A resource for semi-automated, large-scale meta-analyses of ERP data.