Note
Go to the end to download the full example code
The Corrector class
Here we take a look at multiple comparisons correction in meta-analyses.
from pprint import pprint
import matplotlib.pyplot as plt
import seaborn as sns
from nilearn.plotting import plot_stat_map
Download data
from nimare.extract import download_nidm_pain
dset_dir = download_nidm_pain()
Load Dataset
import os
from nimare.dataset import Dataset
from nimare.utils import get_resource_path
dset_file = os.path.join(get_resource_path(), "nidm_pain_dset.json")
dset = Dataset(dset_file)
dset.update_path(dset_dir)
mask_img = dset.masker.mask_img
Multiple comparisons correction in coordinate-based meta-analyses
Tip
For more information multiple comparisons correction and CBMA in NiMARE, see Multiple comparisons correction.
from nimare.meta.cbma.ale import ALE
# First, we need to fit the Estimator to the Dataset.
meta = ALE(null_method="approximate")
results = meta.fit(dset)
# We can check which FWE correction methods are available for the ALE Estimator
# with the ``inspect`` class method.
from nimare.correct import FWECorrector
print(FWECorrector.inspect(results))
['bonferroni', 'montecarlo']
Apply the Corrector to the MetaResult
Now that we know what FWE correction methods are available, we can use one.
The “montecarlo” method is a special one that is implemented within the Estimator, rather than in the Corrector.
corr = FWECorrector(method="montecarlo", n_iters=50, n_cores=2)
cres = corr.transform(results)
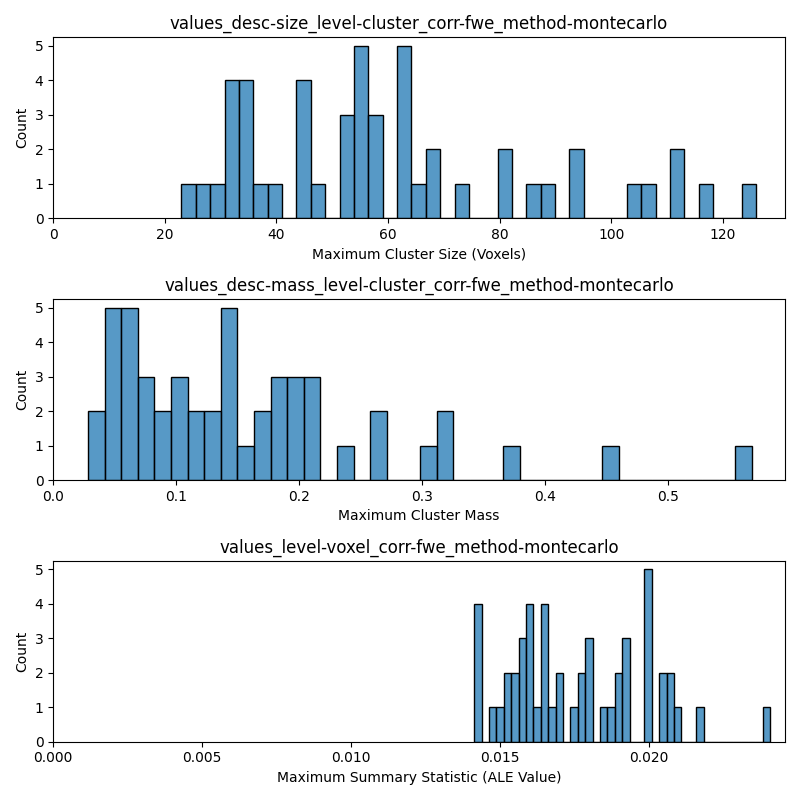
DISTS_TO_PLOT = [
"values_desc-size_level-cluster_corr-fwe_method-montecarlo",
"values_desc-mass_level-cluster_corr-fwe_method-montecarlo",
"values_level-voxel_corr-fwe_method-montecarlo",
]
XLABELS = [
"Maximum Cluster Size (Voxels)",
"Maximum Cluster Mass",
"Maximum Summary Statistic (ALE Value)",
]
fig, axes = plt.subplots(figsize=(8, 8), nrows=3)
null_dists = cres.estimator.null_distributions_
for i_ax, dist_name in enumerate(DISTS_TO_PLOT):
xlabel = XLABELS[i_ax]
sns.histplot(x=null_dists[dist_name], bins=40, ax=axes[i_ax])
axes[i_ax].set_title(dist_name)
axes[i_ax].set_xlabel(xlabel)
axes[i_ax].set_xlim(0, None)
fig.tight_layout()
0%| | 0/50 [00:00<?, ?it/s]
2%|▏ | 1/50 [00:07<05:57, 7.29s/it]
6%|▌ | 3/50 [00:08<01:48, 2.30s/it]
10%|█ | 5/50 [00:09<01:02, 1.39s/it]
14%|█▍ | 7/50 [00:10<00:43, 1.01s/it]
16%|█▌ | 8/50 [00:10<00:34, 1.22it/s]
18%|█▊ | 9/50 [00:11<00:33, 1.22it/s]
20%|██ | 10/50 [00:11<00:27, 1.46it/s]
22%|██▏ | 11/50 [00:12<00:28, 1.38it/s]
24%|██▍ | 12/50 [00:12<00:21, 1.74it/s]
26%|██▌ | 13/50 [00:13<00:23, 1.59it/s]
28%|██▊ | 14/50 [00:13<00:20, 1.79it/s]
30%|███ | 15/50 [00:14<00:21, 1.66it/s]
32%|███▏ | 16/50 [00:14<00:17, 1.92it/s]
34%|███▍ | 17/50 [00:15<00:18, 1.81it/s]
36%|███▌ | 18/50 [00:16<00:17, 1.87it/s]
38%|███▊ | 19/50 [00:16<00:17, 1.80it/s]
40%|████ | 20/50 [00:17<00:15, 1.99it/s]
42%|████▏ | 21/50 [00:17<00:15, 1.88it/s]
44%|████▍ | 22/50 [00:18<00:15, 1.84it/s]
46%|████▌ | 23/50 [00:18<00:14, 1.85it/s]
48%|████▊ | 24/50 [00:19<00:14, 1.77it/s]
50%|█████ | 25/50 [00:19<00:12, 2.05it/s]
52%|█████▏ | 26/50 [00:20<00:13, 1.72it/s]
54%|█████▍ | 27/50 [00:20<00:11, 1.99it/s]
56%|█████▌ | 28/50 [00:21<00:12, 1.79it/s]
58%|█████▊ | 29/50 [00:21<00:09, 2.13it/s]
60%|██████ | 30/50 [00:22<00:11, 1.67it/s]
62%|██████▏ | 31/50 [00:22<00:09, 2.07it/s]
64%|██████▍ | 32/50 [00:23<00:11, 1.57it/s]
68%|██████▊ | 34/50 [00:24<00:09, 1.67it/s]
72%|███████▏ | 36/50 [00:25<00:07, 1.80it/s]
76%|███████▌ | 38/50 [00:27<00:06, 1.80it/s]
80%|████████ | 40/50 [00:28<00:05, 1.88it/s]
82%|████████▏ | 41/50 [00:28<00:04, 2.06it/s]
84%|████████▍ | 42/50 [00:29<00:04, 1.82it/s]
86%|████████▌ | 43/50 [00:29<00:03, 1.93it/s]
88%|████████▊ | 44/50 [00:30<00:03, 1.92it/s]
90%|█████████ | 45/50 [00:30<00:02, 1.86it/s]
92%|█████████▏| 46/50 [00:31<00:02, 1.86it/s]
94%|█████████▍| 47/50 [00:31<00:01, 1.93it/s]
96%|█████████▌| 48/50 [00:32<00:01, 1.98it/s]
98%|█████████▊| 49/50 [00:32<00:00, 1.84it/s]
100%|██████████| 50/50 [00:33<00:00, 2.01it/s]
100%|██████████| 50/50 [00:33<00:00, 1.51it/s]
You can also look at the description of the Corrector.
print("Description:")
pprint(cres.description_)
print("References:")
pprint(cres.bibtex_)
Description:
('An activation likelihood estimation (ALE) meta-analysis '
'\\citep{turkeltaub2002meta,turkeltaub2012minimizing,eickhoff2012activation} '
'was performed with NiMARE 0.2.2+2.gbf7df33.dirty (RRID:SCR_017398; '
'\\citealt{Salo2023}), using a(n) ALE kernel. An ALE kernel '
'\\citep{eickhoff2012activation} was used to generate study-wise modeled '
'activation maps from coordinates. In this kernel method, each coordinate is '
'convolved with a Gaussian kernel with full-width at half max values '
'determined on a study-wise basis based on the study sample sizes according '
'to the formulae provided in \\cite{eickhoff2012activation}. For voxels with '
'overlapping kernels, the maximum value was retained. ALE values were '
'converted to p-values using an approximate null distribution '
'\\citep{eickhoff2012activation}. The input dataset included 267 foci from 21 '
'experiments, with a total of 334 participants. Family-wise error rate '
'correction was performed using a Monte Carlo procedure. In this procedure, '
'null datasets are generated in which dataset coordinates are substituted '
'with coordinates randomly drawn from the meta-analysis mask, and maximum '
'values are retained. This procedure was repeated 50 times to build null '
'distributions of summary statistics, cluster sizes, and cluster masses. '
'Clusters for cluster-level correction were defined using edge-wise '
'connectivity and a voxel-level threshold of p < 0.001 from the uncorrected '
'null distribution.')
References:
('@article{Salo2023,\n'
' doi = {10.52294/001c.87681},\n'
' url = {https://doi.org/10.52294/001c.87681},\n'
' year = {2023},\n'
' volume = {3},\n'
' pages = {1 - 32},\n'
' author = {Taylor Salo and Tal Yarkoni and Thomas E. Nichols and '
'Jean-Baptiste Poline and Murat Bilgel and Katherine L. Bottenhorn and Dorota '
'Jarecka and James D. Kent and Adam Kimbler and Dylan M. Nielson and Kendra '
'M. Oudyk and Julio A. Peraza and Alexandre Pérez and Puck C. Reeders and '
'Julio A. Yanes and Angela R. Laird},\n'
' title = {NiMARE: Neuroimaging Meta-Analysis Research Environment},\n'
' journal = {Aperture Neuro}\n'
'}\n'
'@article{eickhoff2012activation,\n'
' title={Activation likelihood estimation meta-analysis revisited},\n'
' author={Eickhoff, Simon B and Bzdok, Danilo and Laird, Angela R and Kurth, '
'Florian and Fox, Peter T},\n'
' journal={Neuroimage},\n'
' volume={59},\n'
' number={3},\n'
' pages={2349--2361},\n'
' year={2012},\n'
' publisher={Elsevier},\n'
' url={https://doi.org/10.1016/j.neuroimage.2011.09.017},\n'
' doi={10.1016/j.neuroimage.2011.09.017}\n'
'}\n'
'@article{turkeltaub2002meta,\n'
' title={Meta-analysis of the functional neuroanatomy of single-word '
'reading: method and validation},\n'
' author={Turkeltaub, Peter E and Eden, Guinevere F and Jones, Karen M and '
'Zeffiro, Thomas A},\n'
' journal={Neuroimage},\n'
' volume={16},\n'
' number={3},\n'
' pages={765--780},\n'
' year={2002},\n'
' publisher={Elsevier},\n'
' url={https://doi.org/10.1006/nimg.2002.1131},\n'
' doi={10.1006/nimg.2002.1131}\n'
'}\n'
'@article{turkeltaub2012minimizing,\n'
' title={Minimizing within-experiment and within-group effects in activation '
'likelihood estimation meta-analyses},\n'
' author={Turkeltaub, Peter E and Eickhoff, Simon B and Laird, Angela R and '
'Fox, Mick and Wiener, Martin and Fox, Peter},\n'
' journal={Human brain mapping},\n'
' volume={33},\n'
' number={1},\n'
' pages={1--13},\n'
' year={2012},\n'
' publisher={Wiley Online Library},\n'
' url={https://doi.org/10.1002/hbm.21186},\n'
' doi={10.1002/hbm.21186}\n'
'}')
Show corrected results
MAPS_TO_PLOT = [
"z",
"z_desc-size_level-cluster_corr-FWE_method-montecarlo",
"z_desc-mass_level-cluster_corr-FWE_method-montecarlo",
"z_level-voxel_corr-FWE_method-montecarlo",
]
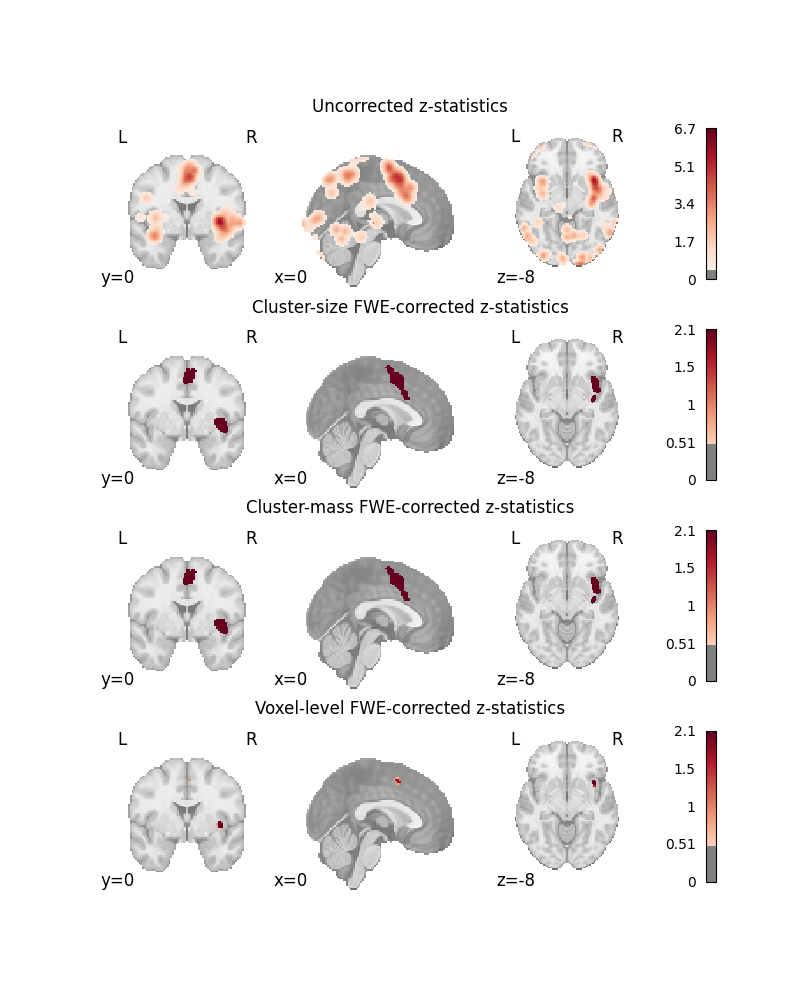
TITLES = [
"Uncorrected z-statistics",
"Cluster-size FWE-corrected z-statistics",
"Cluster-mass FWE-corrected z-statistics",
"Voxel-level FWE-corrected z-statistics",
]
fig, axes = plt.subplots(figsize=(8, 10), nrows=4)
for i_ax, map_name in enumerate(MAPS_TO_PLOT):
title = TITLES[i_ax]
plot_stat_map(
cres.get_map(map_name),
draw_cross=False,
cmap="RdBu_r",
threshold=0.5,
cut_coords=[0, 0, -8],
figure=fig,
axes=axes[i_ax],
)
axes[i_ax].set_title(title)
Multiple comparisons correction in image-based meta-analyses
from nimare.correct import FDRCorrector
from nimare.meta.ibma import Stouffers
meta = Stouffers(resample=True)
results = meta.fit(dset)
print(f"FWECorrector options: {FWECorrector.inspect(results)}")
print(f"FDRCorrector options: {FDRCorrector.inspect(results)}")
FWECorrector options: ['bonferroni']
FDRCorrector options: ['indep', 'negcorr']
Note that the FWECorrector does not support a “montecarlo” method for the Stouffers Estimator. This is because NiMARE does not have a Monte Carlo-based method implemented for most IBMA algorithms.
Apply the Corrector to the MetaResult
corr = FDRCorrector(method="indep", alpha=0.05)
cres = corr.transform(results)
Show corrected results
fig, axes = plt.subplots(figsize=(8, 6), nrows=2)
plot_stat_map(
cres.get_map("z"),
draw_cross=False,
cmap="RdBu_r",
threshold=0.5,
cut_coords=[0, 0, -8],
figure=fig,
axes=axes[0],
)
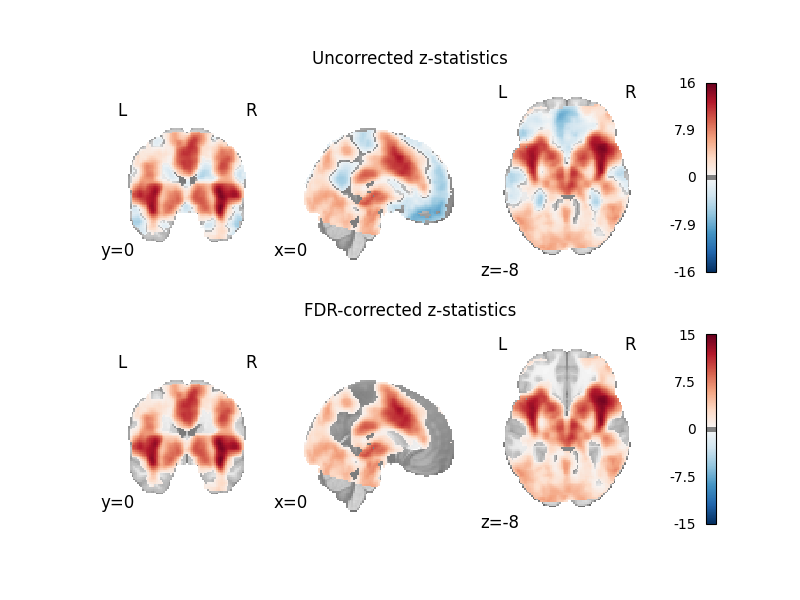
axes[0].set_title("Uncorrected z-statistics")
plot_stat_map(
cres.get_map("z_corr-FDR_method-indep"),
draw_cross=False,
cmap="RdBu_r",
threshold=0.5,
cut_coords=[0, 0, -8],
figure=fig,
axes=axes[1],
)
axes[1].set_title("FDR-corrected z-statistics")
/home/docs/checkouts/readthedocs.org/user_builds/nimare/envs/latest/lib/python3.9/site-packages/nilearn/_utils/niimg.py:61: UserWarning: Non-finite values detected. These values will be replaced with zeros.
warn(
Text(0.5, 1.0, 'FDR-corrected z-statistics')
Total running time of the script: (0 minutes 54.344 seconds)