Note
The NiMARE NeuroLibre preprint is a valuable resource that provides examples on how to utilize the package. However, it is important to note that this publication is static and will not be updated as NiMARE evolves. Despite potential changes to the syntax of the code, the publication remains an excellent reference to understand the various applications of NiMARE.
Examples
Working with datasets
NiMARE stores meta-analytic data in its Dataset class.
Dataset objects may contain a range of elements, including coordinates (for coordinate-based meta-analysis),
links to statistical maps (for image-based meta-analysis), article text, label weights, and other metadata.
Additionally, NiMARE contains fetching and conversion tools for a number of meta-analytic resources, including Neurosynth, NeuroQuery, NeuroVault, and, to a limited extent, BrainMap. In the examples below, we show what a Dataset can do and exhibit tools for working with data from external meta-analytic resources.
Performing meta-analyses
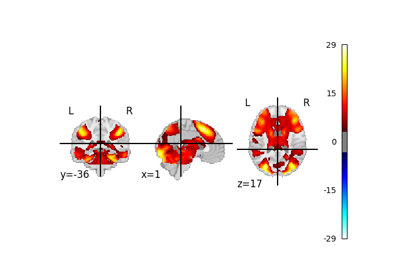
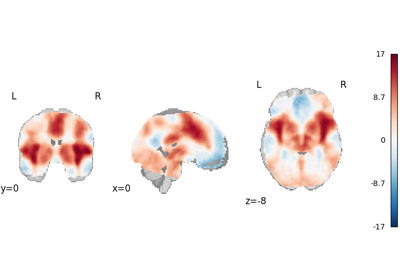
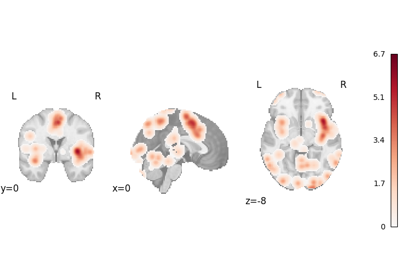
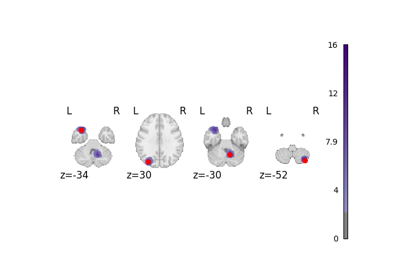
NiMARE implements a number of coordinate- and image-based meta-analysis algorithms in its meta module.
In the examples below, we exhibit a range of meta-analyses that can be done with coordinates and/or images in NiMARE.
For more information about the components that go into coordinate-based meta-analyses in NiMARE, see Coordinate-based meta-analysis in NiMARE, as well as Outputs of NiMARE.
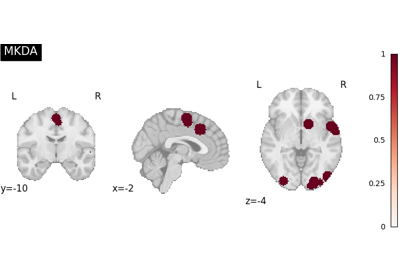
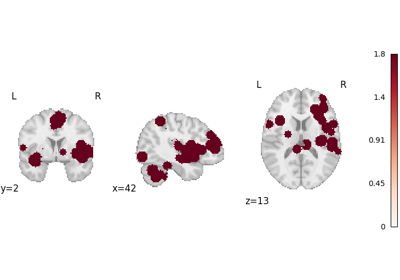
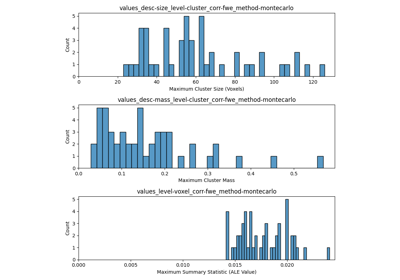
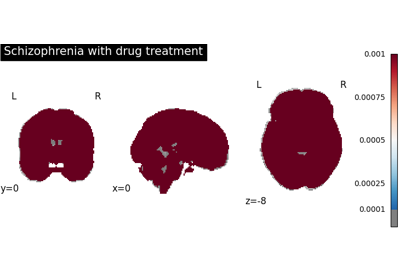
Run a coordinate-based meta-analysis (CBMA) workflow
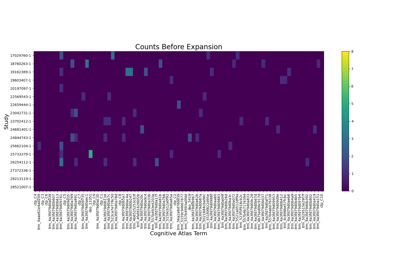
Annotating Datasets
Annotation tools within NiMARE (annotate) refer to methods which
assign labels to studies in a Dataset, generally based on study text.
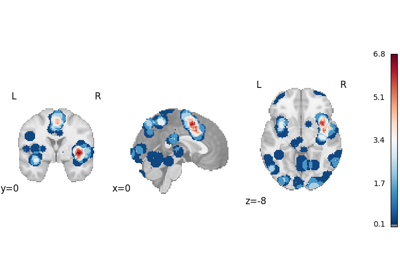
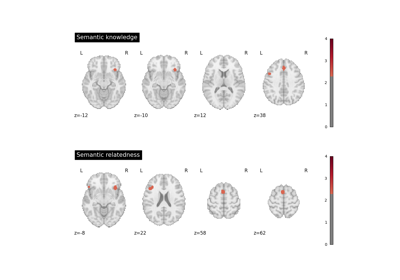
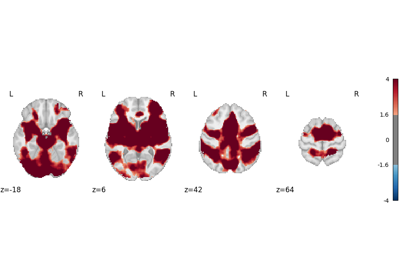
Decoding ROIs and images
Functional characterization analysis refers to methods which use meta-analytic databases to characterize, or “decode”, brain regions or statistical maps in terms of tasks and/or mental processes. For more information about functional characterization analysis, see Meta-analytic functional decoding.